Sampling the Neighborhoods of the Gut Microbiome
In recent years, changes in the gut microbiome have been linked to many health conditions, including obesity, GI disorders, cancer, and even depression.
But the gut microbiome–composed of hundreds of different species of bacteria–is a complex community and a challenge for scientists to unravel.
One big challenge is the spatial distribution of different microbes, which are not evenly distributed throughout the gut. The gut microbiome is like a large city, with multiple neighborhoods, each with its own mix of occupants and features.
A new method developed by researchers at Columbia University Vagelos College of Physicians and Surgeons should help scientists locate and characterize these neighborhoods, which could shed light on how microbes influence the health of their hosts.
Homogeneous samples are out
Existing techniques are not up to the task of probing the neighborhoods of the microbiome: Techniques that can identify all species in the gut microbiome only work with homogenized samples (like stool), but methods that preserve spatial information can only cope with a handful of species.
In devising a better method, Harris Wang, PhD, and graduate student Ravi Sheth in the Department of Systems Biology took inspiration from the way plant ecologists use plot sampling to survey sites.
Starting with small tissue samples from the GI tracts of mice, Wang and Sheth added gel to fix the bacteria in place. After freezing the gel, they broke the tissue samples into tiny particles.
“Each small particle is essentially like a quadrant that a plant ecologist would place in a forest,” Sheth says. “The particle preserves the species that exist in a particular neighborhood.”
The researchers then took advantage of new, high-throughput techniques to process the data and identify all the species present on each separate particle.
Change induced by high-fat diet

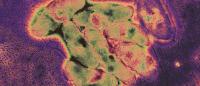
Microbial neighborhoods in the mouse gut change after a switch from a low-fat to a high-fat diet. Green dots represent a distinct microbial community with a low-fat diet; orange dots represent microbial communities with a high-fat diet. Only a few communities were observed in both diets. Image: Harris Wang and Ravi Sheth / Columbia University Irving Medical Center.
Wang and Sheth tested the technique with mice who switched from a low-fat to a high-fat diet.
Diet is known to change the abundance of specific bacteria in the gut within days, but the new technique also revealed that the switch caused wholesale changes of microbial neighborhoods.
“Specific regions of bacteria were entirely lost with a switch in diet,” Sheth says. “This was exciting to us as it will give us clues to understanding how that change happens and how the change may impact health.”
The method is currently designed to be used in mice, but Sheth and Wang are working to adapt the technique for use in people.
References
More Information
Harris Wang, PhD, is assistant professor of systems biology at Columbia University Vagelos College of Physicians and Surgeons. He is also a member of the Department of Pathology & Cell Biology, the JP Sulzberger Columbia Genome Center, and the Center for Cancer Systems Therapeutics at Columbia University Irving Medical Center.
Ravi Sheth is a graduate student in the Integrated Program in Cellular, Molecular and Biomedical Studies at Columbia University.
The research, titled “Spatial metagenomic characterization of microbial biogeography in the gut,” was published July 22 in Nature Biotechnology.
Other authors: Mingqiang Li (Columbia University), Weiqian Jiang (Columbia University), Peter A. Sims (Columbia University Irving Medical Center), and Kam W. Leong (Columbia University Irving Medical Center).
The research was supported by grants from the NIH (1R01AI132403, 1R01DK118044, R01GM110494, and K01EB016071), Office of Naval Research (N00014-15-1-2704), and Burroughs Welcome Fund PATH (1016691); and fellowships from the Fannie and John Hertz Foundation and the National Science Foundation (DGE-1644869).
Harris Wang and Ravi Sheth are inventors on a provisional patent application filed by the Trustees of Columbia University in the City of New York regarding this work.